Abstract
Objective:To investigate the causal link between natural killer (NK) cell receptor CD244 and head and neck cancer (HNC), and to evaluate sphingomyelin (SM) as a potential mediator of this relationship.
Methods:A two-sample Mendelian randomization (MR) analysis was conducted using genome-wide association study (GWAS) summary statistics from European cohorts (NCD244=14,735; NHNC=2,131; NSM=8,260). Instrumental variables (32 SNPs for CD244, 79 SNPs for HNC) were selected at genome-wide significance (P < 5×10⁻⁸/5×10⁻⁶) after rigorous linkage disequilibrium clumping. Causal estimates were generated using inverse variance weighting (IVW), MR-Egger, and weighted median methods. Mediation analysis quantified SM's contribution to CD244-HNC associations.
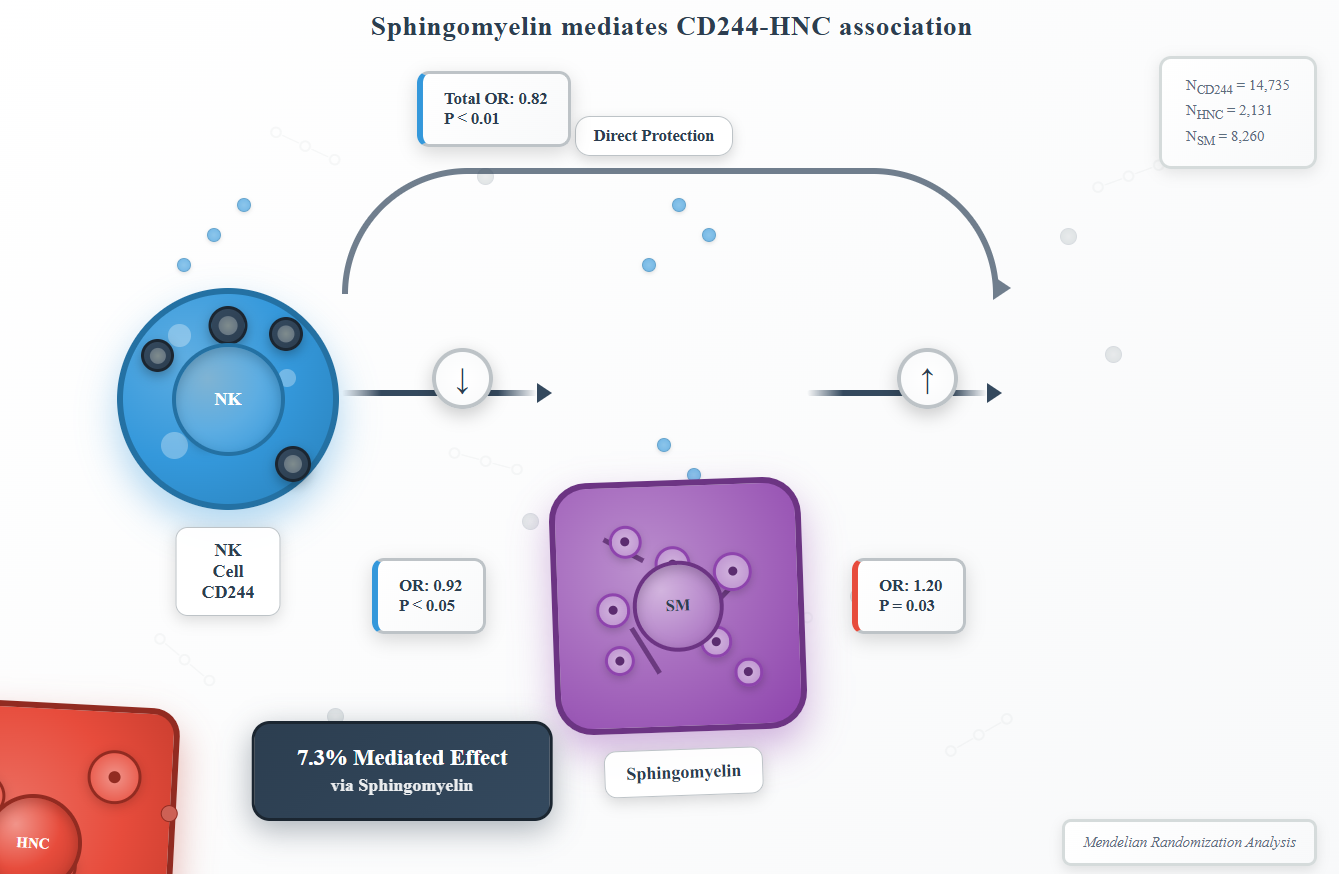
Results:Genetically predicted CD244 levels showed a protective effect against HNC (IVW OR=0.82 per SD, 95% CI:0.71-0.94, P < 0.01), while elevated CD244 was associated with reduced SM levels (IVW OR=0.92, 95% CI:0.86-0.98, P < 0.05). Conversely, SM demonstrated risk-enhancing effects on HNC (IVW OR=1.20, 95% CI:1.01-1.41, P=0.03). Mediation analysis revealed that SM accounted for 7.3% of the total CD244-associated HNC risk.
Conclusion:This study establishes a causal protective role of CD244 in HNC pathogenesis, partially mediated through SM. The findings highlight CD244 and SM metabolism as potential targets for HNC immunomodulatory therapies. Future research should validate these mechanisms in diverse populations and explore additional mediators.
Keywords:NK cell, CD244, Head and Neck Cancer, Sphingomyelin, Mendelian Randomization
Introduction
Natural Killer (NK) cells are a type of cytotoxic lymphocyte that play a significant role in the innate immune system. These cells can recognize and eliminate infected cells as well as cancerous cells without prior sensitization or activation [1] .
The receptor 2B4 (also known as CD244) is a protein found on the surface of NK cells. It belongs to the signaling lymphocytic activation molecule (SLAM) family of receptors [2] . It serves as a co-stimulatory molecule, contributing to NK cell activation and cytotoxicity when engaged [3–5] . The interaction of CD244 with its ligands on target cells or other immune cells helps regulate NK cell-mediated cytotoxicity and can influence the immune response against infections and tumors [6,7] . Dysfunction or alterations in CD244 signaling might impact the activity of NK cells and consequently affect immune responses and disease outcomes [8–10] .
Head and neck carcinoma (HNC) is a type of cancer that primarily affects the oral cavity, sinonasal cavity, pharynx, and larynx [11] . It ranks about sixth in terms of incidence and mortality rate worldwide, with approximately 800,000 new cases and 44,000 deaths reported annually [12] . Current diagnostic methods for HNC include physical examination, endoscopic examination, imaging examination, biopsy, and tumor biomarker detection [13] . However, these methods have their limitations. For example, the sensitivity of visual examination of the mouth for detecting oral precancerous and cancerous lesions varies, and when the affected area is not easily accessible, the accuracy may be lower [14] . This challenge also applies to tissue biopsy. Therefore, there is a need for alternative diagnostic approaches and Therapeutic target.
Sphingomyelin (SM) is a type of lipid molecule that belongs to the class of phospholipids and is an essential component of cell membranes. It consists of a sphingosine backbone linked to phosphorylcholine, forming a molecule that plays a crucial role in the structure and function of cell membranes. It plays significant roles in cell signaling, maintaining membrane integrity, and is particularly important in the nervous system as a major component of neuronal membranes [15] .
Mendelian Randomization (MR) is a causal inference method that utilizes genetic variation as an instrumental variable to assess the impact of exposure factors on outcomes in observational data. This ensures a relatively low correlation between the exposure variable and other potential confounding factors, thereby enhancing the credibility of causal inference. Simultaneously, MR helps to avoid reverse causality, as genetic variations typically precede exposure factors, reducing the likelihood of reverse causation [16] . The study aims to ascertain whether there is a causal relationship between CD244 and HNC, and to evaluate the extent to which the effects of CD244 on HNC are mediated by sphingomyelin.
Methods
Study design
The data utilized in our analysis were sourced from publicly available datasets and received approval from the institutional review committee in the respective studies. Consequently, no additional sanctions were required. All outcomes derived from this analysis are comprehensively outlined in the main article and its supplementary materials.
In this research, we explored the bidirectional causal relationship between CD244 and HNC through a two-sample MR approach. In our investigation, single nucleotide polymorphisms (SNPs) were identified as instrumental variables (IVs) [17] .
GWAS summary data sources
All the data employed in our research were sourced from publicly accessible datasets, and the individuals included in the Genome-Wide Association Study (GWAS) belonged to European ancestry.
The genetic correlations related to CD244 levels were extracted from a GWAS meta-analysis conducted by Zhao and collaborators, encompassing a total of 14,735 participants (https://www.ebi.ac.uk/gwas/studies/GCST90274771) [18] .
We obtained summary statistics for HNC from the FinnGen research project (https://www.finngen.fi/en). The research project conducted a GWAS involving 2,131 individuals of Finnish descent, with a gender distribution of 1,466 males and 665 females [19] .
Sphingomyelin level summary statistics were extracted from a study conducted by Chen and collaborators [20] , involving 8,260 unrelated individuals of European descent. The original publication detailed the study design, encompassing sample collection, quality control protocols, and imputation techniques. The GWAS data originated from separate consortia or organizations, ensuring the absence of sample overlap.
Instrumental variable selection and data harmonization
We incorporated SNPs that achieved genome-wide significance (P < 5 × 10-8). In cases where no SNPs met the criteria for genome-wide significance as instrumental variables (IVs), SNPs with significance levels below the genome-wide threshold (P < 5 × 10-6) were considered as potential IVs. Subsequently, these SNPs underwent clustering based on linkage disequilibrium (with a window size of 10,000 kb and r2 < 0.001). The estimation of linkage disequilibrium levels was derived from European samples within the 1000 Genomes Project [21] . If a particular SNP associated with the exposure was not present in the outcome dataset, proxy SNPs were employed using LD tagging. Palindromic and ambiguous SNPs were excluded from the instrumental variables utilized in the Mendelian randomization analysis [22] .
Statistical analysis
We conducted MR analysis using the R software (version 4.2.0, http://www.r-project.org).
To assess the causal relationship between CD244 levels and HNC, we employed various statistical methods, including inverse variance weighting (IVW) [23] , weighted median-based [24] and mode-based methods [25] . These analyses were predominantly conducted using the 'Mendelian-Randomization' package (version 0.4.3) [26] .
Cochran’s Q statistic and corresponding p values were employed to examine the heterogeneity among the chosen instrumental variables (IVs). In case the null hypothesis is rejected, the random effects Inverse Variance Weighting (IVW) method was employed instead of the fixed-effects IVW [23] . To address the influence of horizontal pleiotropy, a widely used approach known as MR-Egger was employed. The utilization of MR-Egger indicates the potential existence of horizontal multiplicity, particularly if the intercept term is found to be significant [27] . Furthermore, we employed the MR-PRESSO method, a resilient approach, to detect and remove potential horizontal pleiotropic outliers that could have a substantial impact on the estimation results within the MR-PRESSO package [28] . In addition, we utilized scatter plots and funnel plots in our analysis. The scatter plots demonstrated the resilience of outcomes to outliers, while the funnel plots illustrated the robustness of the correlation and the absence of heterogeneity.
Primary analysis
Figure 1 Illustrations in this study depict the associations under investigation. In (A), the total effect between CD244 and head and neck carcinoma (HNC) are presented. "c" represents the total effect utilizing genetically predicted CD244 as the exposure and HNC as the outcome, while "d" depicts the total effect using genetically predicted HNC as the exposure and CD244 as the outcome. Moving to (B), the total effect is further dissected into two components: (i) the indirect effect employing a two-step approach (where "a" denotes the total effect of CD244 on SM, and "b" signifies the effect of SM on HNC) and the product method (a × b), and (ii) the direct effect (c′ = c – a × b). The proportion mediated is calculated by dividing the indirect effect by the total effect.
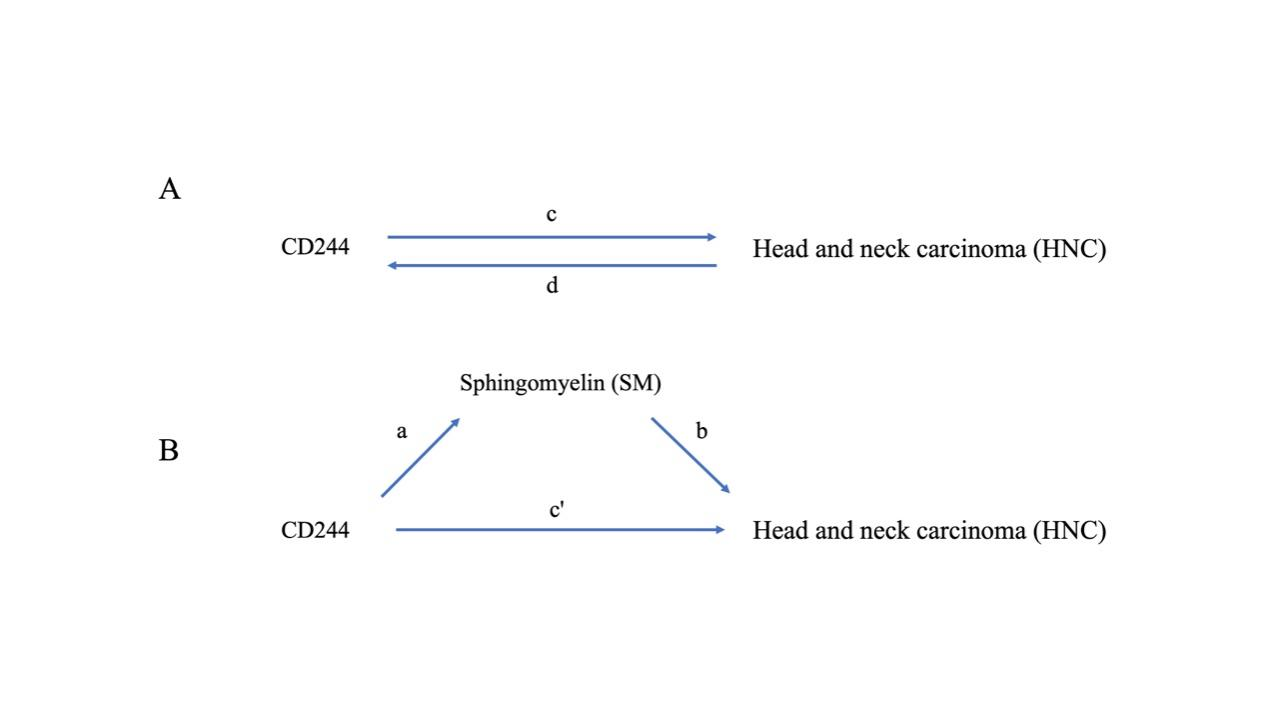
Figure 1 provides a schematic overview of the analysis. A two-sample bidirectional MR was conducted to examine the reciprocal causation between CD244 levels and HNC, as illustrated in Figure 1A. This analysis focused on determining the total effects.
Inverse variance weighting (IVW) employs meta-analysis to amalgamate the Wald ratios of causal effects associated with each individual SNP [22,29] . Then, MR-Egger [27] and weighted-median [24] methods were employed as supplementary analyses to complement the Inverse Variance Weighting (IVW) method. Various methods, aligned with different validity assumptions, have been employed to derive MR estimates. The utilization of the Inverse Variance Weighting (IVW) method is contingent upon the assumption that all Single Nucleotide Polymorphisms (SNPs) serve as valid instrumental variables. Hence, this approach has the potential to generate precise estimation outcomes. MR-Egger evaluates directional pleiotropy concerning instrumental variables, with the intercept serving as an estimate of the average pleiotropy associated with genetic variation. The weighted median method offers the benefit of retaining greater precision, indicated by a smaller standard deviation, in comparison to MR-Egger analysis. In situations involving horizontal pleiotropy, the weighted median consistently yields estimates, maintaining accuracy even when 50% of the genetic variants are deemed invalid instrumental variables [30] .
Mediation analysis
We performed a mediation analysis using a two-step MR design to explore the potential mediation role of sphingomyelin levels in the causal pathway from CD244 levels to the occurrence of HNC (Figure 1B). The total effect can be broken down into an indirect effect, which operates through mediators, and a direct effect, which occurs independently of any mediators [31] . The overall influence of CD244 levels on HNC was dissected into two components: 1) the direct effects of CD244 levels on HNC (represented as c' in Figure 1B) and 2) the indirect effects mediated by CD244 through an intermediary (expressed as a × b in Figure 1B). The proportion mediated by the intermediary effect was determined by dividing the indirect effect by the total effect.
Results
Association of CD244 levels with HNC
After the exclusion of palindromic and ambiguous SNPs, along with SNPs lacking proxies and those exhibiting incorrect causal directions flagged by MR Steiger filtering, a total of 32 SNPs were found to be linked with CD244 levels. Additionally, 79 SNPs associated with HNC were identified and utilized as instrumental variables (refer to Supplementary Tables S1).
Figure 2 Forest plot to visualize causal effect of each single SNP on total HNC risk.
Figure 2 Forest plot to visualize causal effect of each single SNP on total HNC risk.
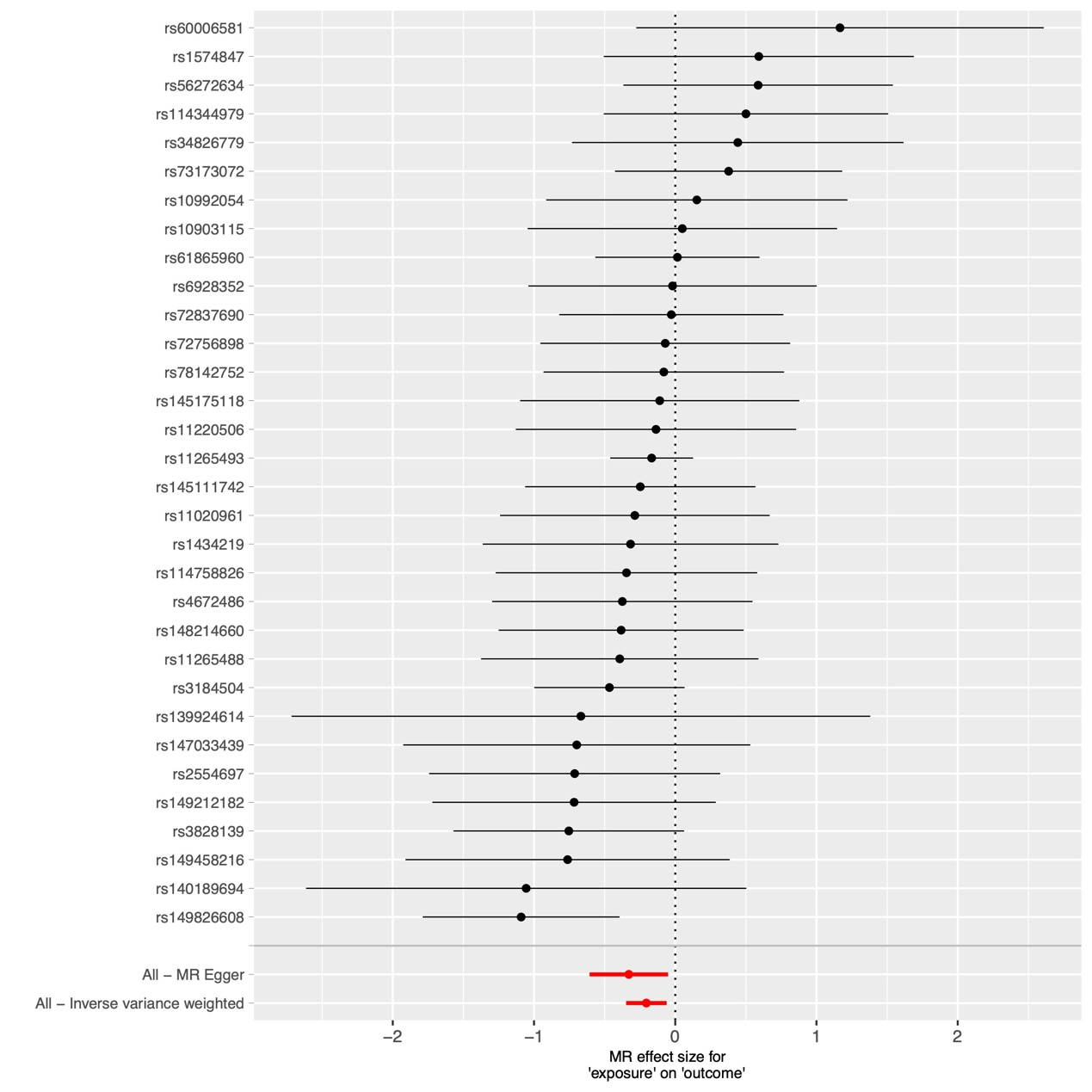
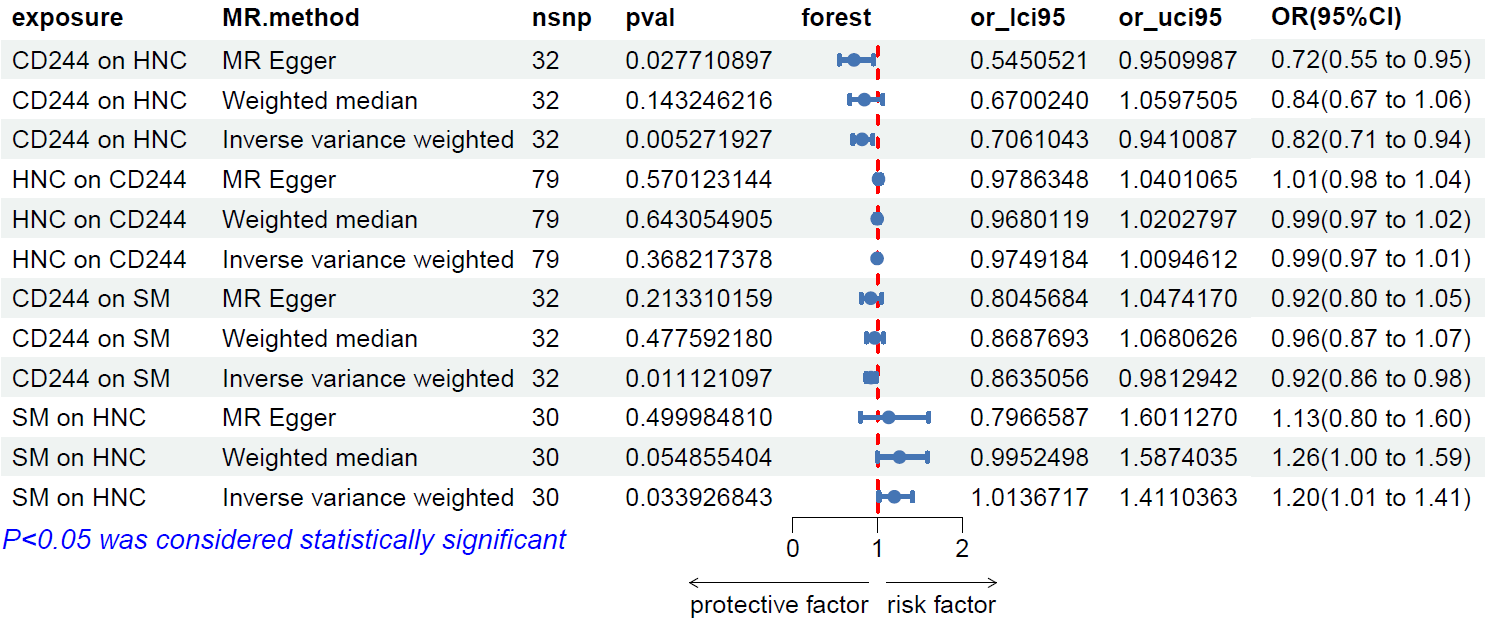
We utilized IVW, MR-Egger, and weighted median regression techniques to estimate the causal association between CD244 levels and HNC, as illustrated in Figures 2 and 3. Among the three statistical methods, two of them supported the negative association of CD244 levels with HNC (IVW odds ratio [OR] per SD increase in CD244 levels = 0.82 [95% CI,0.71-0.94] , P < 0.01; MR-Egger OR per SD increase in CD244 levels = 0.72 [95% CI,0.55-0.95] , P < 0.05). Nevertheless, our MR analysis did not uncover any indications of reverse causation concerning genetically predicted CD244 levels and HNC. In other words, there is no evidence suggesting a causal relationship where HNC influences CD244 levels. The odds ratio (OR) calculated through the Inverse Variance Weighting (IVW) method was 0.99 [95% CI,0.97-1.01;p=0.37] , as illustrated in Figure 3.
Figure 3 Forest plot to visualize the causal effects of SM with CD244 and HNC.
Association of CD244 levels with SM
We successfully identified 32 instrumental variables with genome-wide significance. This accomplishment involved the exclusion of palindromic and ambiguous SNPs, along with SNPs lacking proxies and those flagged by MR Steiger filtering for being in the incorrect causal direction. Based on the analyses conducted using the IVW, MR–Egger, and weighted median methods, it was observed that CD244 levels exhibited a negative correlation with the risk of sphingomyelin (IVW method, odds ratio 0.92; [95% CI,0.86-0.98] , P < 0.05). The findings are visually represented in Figure 3.
Association of sphingomyelin with HNC
We have identified and presented all genetic instruments linked to SM at the genome-wide significance level (P < 5 x 10-8). In Figure 3, the genetically predicted levels of sphingomyelin showed a positive correlation with HNC [OR=1.20,95%CI1.01-1.41;P=0.03] when utilizing the IVW method.
Proportion of the association between CD244 levels and HNC mediated by SM
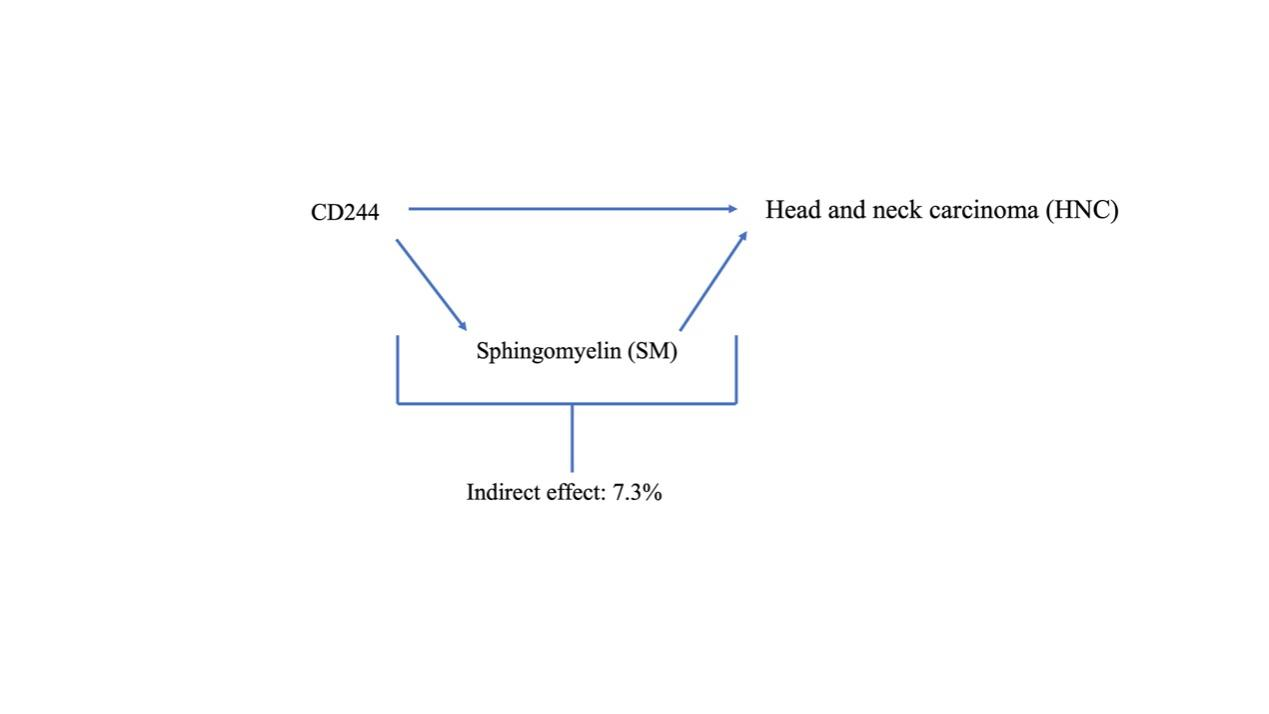
We conducted an analysis of SM as a mediator in the pathway connecting CD244 levels to HNC. The results of our study indicate a correlation between elevated CD244 levels and decreased SM, thereby leading to an increased risk of HNC. As depicted in Figure 4, our research illustrates that sphingomyelin accounted for 7.3% of the elevated HNC risk associated with CD244 levels.
Figure 4 Schematic diagram of the SM mediation effect.
Sensitivity analysis
Several sensitivity analyses were implemented to explore and address the potential presence of pleiotropy in the causal estimates. The absence of heterogeneity or asymmetry among these SNPs in their causal relationship was revealed through Cochran’s Q-test and funnel plot, as indicated in Supplementary Table S2 and Supplementary Figure S1. Our study utilized the MR-Egger intercept, indicating no signs of pleiotropy at the directional level of the CD244 levels instrument, as presented in Supplementary Table S2. Horizontal pleiotropy was not observed based on the MR-PRESSO global test, as detailed in Supplementary Table S2. The impact of each SNP on the overall causal estimates was validated through leave-one-out analysis, as depicted in Supplementary Figure S2. Following the exclusion of each SNP, a comprehensive reevaluation of the MR analysis for the remaining SNPs was undertaken. The results consistently upheld the significance of the causal relationship, underscoring the pivotal role of all SNPs in establishing its importance.
Discussion
Recent studies [32,33] have explored the correlation between CD244 and HNC. However, existing evidence relies predominantly on observational studies, potentially susceptible to the influence of confounding factors. Our study sought to elucidate the causal relationship between CD244 and HNC. Employing MR analysis, we examined the link between CD244 and HNC using available GWAS data, aiming to determine whether the observed causal relationship involves mediation through SM. Our findings indicate that CD244 is associated with a reduced risk of HNC (an 18% decrease in HNC risk for every 1 standard deviation increase in CD244). Furthermore, our analysis suggests that 7.3% of this effect is mediated through SM.
We are pioneering the exploration of the causal association between CD244 and the susceptibility to HNC through MR techniques. Additionally, we provide evidence showcasing SM as a mediator in this relationship. Our results align with previous observations derived from conventional observational approaches. As part of the SLAM family, 2B4, when binding with its ligand CD48, exhibits dual functionalities involving both activation and inhibition. Notably, 2B4 plays a crucial regulatory role in conditions such as malignant tumors, chronic viral infections, and autoimmune disorders [34] . Altavater et al. discovered that the 2B4 receptor plays a powerful role as a costimulatory factor in NK cells. By integrating the internal domain of 2B4 into chimeric receptors with T-cell receptor zeta, they observed a substantial improvement in various facets of NK cell activation when encountering leukemia or neuroblastoma cells expressing antigens. This improvement includes heightened lysosome release, enhanced growth inhibition, and the successful overcoming of NK cell resistance in autologous leukemia cells, all while preserving antigen specificity [35] . In contrast, Agresta et al. observed a correlation between elevated CD244 expression and PD-L1 expression, as well as heightened spontaneous expression of immune-suppressive mediators, in a study focused on head and neck squamous cell cancer (HNSCC) [36] . Moreover, their findings indicated that mice lacking CD244 (CD244-/-) exhibited significantly impaired tumor growth in the context of HNSCC. When wild-type (WT) mice were subjected to interventional treatment with an anti-CD244 monoclonal antibody, there was a notable reduction in the growth of established HNC tumors, accompanied by an increase in tumor-infiltrating CD8+ T cells.
The findings from our analysis demonstrate a positive association between SM and HNC. This conclusion is substantiated by a study conducted by Wang et al. [37] . The correlation between disease progression and the concentrations of SM 42:2 and SM 42:3 was identified, suggesting their potential utility as biomarkers for diagnosing laryngeal cancer. In contrast, Zheng et al. observed a distinctive phenotype in intratumoral NK cells characterized by a smooth and rounded appearance, deviating from the typical villi-rich, rough surface of normal NK cells. This unique phenotype was associated with a specific deficiency in membrane sphingomyelin lipids, leading to an impairment in the ability to kill tumor cells [38] .
The SLAM-family receptors, including CD244, have a substantial impact on various biological processes, such as cytotoxicity, humoral immune responses, autoimmune diseases, lymphocyte development, cell survival, and cell adhesion. Growing evidence suggests their involvement in cancer progression, positioning them as emerging immune checkpoints on T cells [39] . Additionally, the SM metabolic pathway has gained recognition as a novel therapeutic target. A recent study demonstrated the use of anti-angiogenic drugs to enhance chemotherapy sensitivity in tumors by specifically targeting SM [40] .
This study is subject to several limitations. Firstly, the analysis was conducted using data exclusively from the European population, which restricts the generalizability of our findings. Secondly, the limited number of HNC cases in the GWAS dataset presents a constraint; we anticipate that larger GWAS datasets will become available for future validation. Thirdly, despite efforts to identify and eliminate outlier variants, we cannot entirely rule out the potential impact of horizontal pleiotropy on our results. Fourthly, our study relied on summary-level statistics rather than individual-level data, preventing further exploration of causal relationships within subgroups, such as females and males. Additionally, our findings suggest a relatively low genetic prediction of CD244 mediated by SM, standing at only 7.3%. Consequently, further research is imperative to quantify additional mediators.
Conclusion
In summary, our research established a causal link between CD244 and HNC. While a minor portion of the impact is mediated by SM, the predominant influence of CD244 on HNC remains ambiguous. Additional investigations are warranted to delve into other potential risk factors that may act as mediators in this relationship.
Abbreviations
|
Abbreviation |
Full Term |
|
NK |
Natural Killer cells |
|
CD244 |
Cluster of Differentiation 244 (2B4) |
|
SLAM |
Signaling lymphocytic activation molecule |
|
HNC |
Head and neck cancer |
|
HNSCC |
Head and neck squamous cell carcinoma |
|
SM |
Sphingomyelin |
|
MR |
Mendelian randomization |
|
GWAS |
Genome-wide association study |
|
SNP |
Single nucleotide polymorphism |
|
IVW |
Inverse variance weighting |
|
IVs |
Instrumental variables |
|
OR |
Odds ratio |
|
CI |
Confidence interval |
|
WT |
Wild-type |
Acknowledgements
I would like to thank Dr. Shaokun Liu for his guidance and support throughout this research.
Author contributions
BH and SL conceived and designed the study. BH analyzed data and wrote the manuscript. SL reviewed the paper. Both authors read and approved the final manuscript.
Funding information
None.
Competing Interests
The authors declare that they have no existing or potential commercial or financial relationships that could create a conflict of interest at the time of conducting this study.
Data Availability
All data needed to evaluate the conclusions in the paper are present in the paper or the Supplementary Materials. Additional data related to this paper may be requested from the authors.
References
[1] Nersesian S, Carter EB, Lee SN, Westhaver LP, Boudreau JE. Killer instincts: natural killer cells as multifactorial cancer immunotherapy. Front Immunol. 2023 Nov 28;14:1269614. https://doi.org/10.3389/fimmu.2023.1269614.
[2] Mathew SO, Rao KK, Kim JR, Bambard ND, Mathew PA. Functional role of human NK cell receptor 2B4 (CD244) isoforms. Eur J Immunol. 2009 Jun;39(6):1632-41. https://doi.org/10.1002/eji.200838733.
[3] Veillette A, Latour S. The SLAM family of immune-cell receptors. Curr Opin Immunol. 2003 Jun;15(3):277-85. https://doi.org/10.1016/s0952-7915(03)00041-4.
[4] Brown MH, Boles K, van der Merwe PA, Kumar V, Mathew PA, Barclay AN. 2B4, the natural killer and T cell immunoglobulin superfamily surface protein, is a ligand for CD48. J Exp Med. 1998 Dec 7;188(11):2083-90. https://doi.org/10.1084/jem.188.11.2083.
[5] Latchman Y, McKay PF, Reiser H. Identification of the 2B4 molecule as a counter-receptor for CD48. J Immunol. 1998 Dec 1;161(11):5809-12. https://doi.org/10.4049/jimmunol.161.11.5809.
[6] Chuang SS, Kim MH, Johnson LA, Albertsson P, Kitson RP, Nannmark U, et al. 2B4 stimulation of YT cells induces natural killer cell cytolytic function and invasiveness. Immunology. 2000 Jul;100(3):378-83. https://doi.org/10.1046/j.1365-2567.2000.00031.x.
[7] Mathew SO, Kumaresan PR, Lee JK, Huynh VT, Mathew PA. Mutational analysis of the human 2B4 (CD244)/CD48 interaction: Lys68 and Glu70 in the V domain of 2B4 are critical for CD48 binding and functional activation of NK cells. J Immunol. 2005 Jul 15;175(2):1005-13. https://doi.org/10.4049/jimmunol.175.2.1005.
[8] Wu Y, Kuang DM, Pan WD, Wan YL, Lao XM, Wang D, et al. Monocyte/macrophage-elicited natural killer cell dysfunction in hepatocellular carcinoma is mediated by CD48/2B4 interactions. Hepatology. 2013 Mar;57(3):1107-16. https://doi.org/10.1002/hep.26192.
[9] Pende D, Meazza R, Marcenaro S, Aricò M, Bottino C. 2B4 dysfunction in XLP1 NK cells: More than inability to control EBV infection. Clin Immunol. 2019 Jul;204:31-36. https://doi.org/10.1016/j.clim.2018.10.022.
[10] Schlaphoff V, Lunemann S, Suneetha PV, Jaroszewicz J, Grabowski J, Dietz J, et al. Dual function of the NK cell receptor 2B4 (CD244) in the regulation of HCV-specific CD8+ T cells. PLoS Pathog. 2011 May;7(5):e1002045. https://doi.org/10.1371/journal.ppat.1002045.
[11] Chow LQM. Head and Neck Cancer. N Engl J Med. 2020 Jan 2;382(1):60-72. https://doi.org/10.1056/NEJMra1715715.
[12] Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021 May;71(3):209-249. https://doi.org/10.3322/caac.21660.
[13] Rasheduzzaman M, Kulasinghe A, Dolcetti R, Kenny L, Johnson NW, Kolarich D, et al. Protein glycosylation in head and neck cancers: From diagnosis to treatment. Biochim Biophys Acta Rev Cancer. 2020 Dec;1874(2):188422. https://doi.org/10.1016/j.bbcan.2020.188422.
[14] Mehanna H, Paleri V, West CM, Nutting C. Head and neck cancer--Part 1: Epidemiology, presentation, and prevention. BMJ. 2010 Sep 20;341:c4684. https://doi.org/10.1136/bmj.c4684.
[15] Slotte JP. Biological functions of sphingomyelins. Prog Lipid Res. 2013 Oct;52(4):424-37. https://doi.org/10.1016/j.plipres.2013.05.001.
[16] Emdin CA, Khera AV, Kathiresan S. Mendelian Randomization. JAMA. 2017 Nov 21;318(19):1925-1926. https://doi.org/10.1001/jama.2017.17219.
[17] Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet. 2014 Sep 15;23(R1):R89-98. https://doi.org/10.1093/hmg/ddu328.
[18] Zhao JH, Stacey D, Eriksson N, Macdonald-Dunlop E, Hedman ÅK, Kalnapenkis A, et al. Genetics of circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat Immunol. 2023 Sep;24(9):1540-1551. https://doi.org/10.1038/s41590-023-01588-w.
[19] Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM, et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature. 2023 Jan;613(7944):508-518. https://doi.org/10.1038/s41586-022-05473-8.
[20] Chen Y, Lu T, Pettersson-Kymmer U, Stewart ID, Butler-Laporte G, Nakanishi T, et al. Genomic atlas of the plasma metabolome prioritizes metabolites implicated in human diseases. Nat Genet. 2023 Jan;55(1):44-53. https://doi.org/10.1038/s41588-022-01270-1.
[21] 1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, et al. A map of human genome variation from population-scale sequencing. Nature. 2010 Oct 28;467(7319):1061-73. https://doi.org/10.1038/nature09534.
[22] Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018 May 30;7:e34408. https://doi.org/10.7554/eLife.34408.
[23] Burgess S, Small DS, Thompson SG. A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res. 2017 Oct;26(5):2333-2355. https://doi.org/10.1177/0962280215597579.
[24] Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent Estimation in Mendelian Randomization with Some Invalid Instruments Using a Weighted Median Estimator. Genet Epidemiol. 2016 May;40(4):304-14. https://doi.org/10.1002/gepi.21965.
[25] Hartwig FP, Davey Smith G, Bowden J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int J Epidemiol. 2017 Dec 1;46(6):1985-1998. https://doi.org/10.1093/ije/dyx102.
[26] Yavorska OO, Burgess S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol. 2017 Dec 1;46(6):1734-1739. https://doi.org/10.1093/ije/dyx034.
[27] Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol. 2017 May;32(5):377-389. https://doi.org/10.1007/s10654-017-0255-x.
[28] Verbanck M, Chen CY, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018 May;50(5):693-698. https://doi.org/10.1038/s41588-018-0099-7.
[29] Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013 Nov;37(7):658-65. https://doi.org/10.1002/gepi.21758.
[30] Zhang Y, Liu Z, Choudhury T, Cornelis MC, Liu W. Habitual coffee intake and risk for nonalcoholic fatty liver disease: a two-sample Mendelian randomization study. Eur J Nutr. 2021 Jun;60(4):1761-1767. https://doi.org/10.1007/s00394-020-02369-z.
[31] Carter AR, Sanderson E, Hammerton G, Richmond RC, Davey Smith G, Heron J, et al. Mendelian randomisation for mediation analysis: current methods and challenges for implementation. Eur J Epidemiol. 2021 May;36(5):465-478. https://doi.org/10.1007/s10654-021-00757-1.
[32] Gao P, Lu W, Hu S, Zhao K. Differentially Infiltrated Identification of Novel Diagnostic Biomarkers Associated with Immune Infiltration in Nasopharyngeal Carcinoma. Dis Markers. 2022 Nov 17;2022:3934704. https://doi.org/10.1155/2022/3934704.
[33] Tan L, Cheng D, Wen J, Huang K, Zhang Q. Identification of prognostic hypoxia-related genes signature on the tumor microenvironment in esophageal cancer. Math Biosci Eng. 2021 Sep 7;18(6):7743-7758. https://doi.org/10.3934/mbe.2021384.
[34] Chiha J, Mitchell P, Gopinath B, Burlutsky G, Plant A, Kovoor P, et al. Prediction of Coronary Artery Disease Extent and Severity Using Pulse Wave Velocity. PLoS One. 2016 Dec 22;11(12):e0168598. https://doi.org/10.1371/journal.pone.0168598.
[35] Altvater B, Landmeier S, Pscherer S, Temme J, Schweer K, Kailayangiri S, et al. 2B4 (CD244) signaling by recombinant antigen-specific chimeric receptors costimulates natural killer cell activation to leukemia and neuroblastoma cells. Clin Cancer Res. 2009 Aug 1;15(15):4857-66. https://doi.org/10.1158/1078-0432.CCR-08-2810.
[36] Agresta L, Lehn M, Lampe K, Cantrell R, Hennies C, Szabo S, et al. CD244 represents a new therapeutic target in head and neck squamous cell carcinoma. J Immunother Cancer. 2020 Mar;8(1):e000245. https://doi.org/10.1136/jitc-2019-000245.
[37] Wang H, Luo Y, Chen H, Hou H, Hu Q, Ji M. Non-Targeted Serum Lipidomics Analysis and Potential Biomarkers of Laryngeal Cancer Based on UHPLC-QTOF-MS. Metabolites. 2022 Nov 9;12(11):1087. https://doi.org/10.3390/metabo12111087.
[38] Zheng X, Hou Z, Qian Y, Zhang Y, Cui Q, Wang X, et al. Tumors evade immune cytotoxicity by altering the surface topology of NK cells. Nat Immunol. 2023 May;24(5):802-813. https://doi.org/10.1038/s41590-023-01462-9.
[39] Farhangnia P, Ghomi SM, Mollazadehghomi S, Nickho H, Akbarpour M, Delbandi AA. SLAM-family receptors come of age as a potential molecular target in cancer immunotherapy. Front Immunol. 2023 May 11;14:1174138. https://doi.org/10.3389/fimmu.2023.1174138.
[40] Jacobi J, García-Barros M, Rao S, Rotolo JA, Thompson C, Mizrachi A, et al. Targeting acid sphingomyelinase with anti-angiogenic chemotherapy. Cell Signal. 2017 Jan;29:52-61. https://doi.org/10.1016/j.cellsig.2016.09.010.
 Figures
Figures References
References Peer
Peer Information
InformationFigure 1 Illustrations in this study depict the associations under investigation. In (A), the total effect between CD244 and head and neck carcinoma (HNC) are presented. "c" represents the total effect utilizing genetically predicted CD244 as the exposure and HNC as the outcome, while "d" depicts the total effect using genetically predicted HNC as the exposure and CD244 as the outcome. Moving to (B), the total effect is further dissected into two components: (i) the indirect effect employing a two-step approach (where "a" denotes the total effect of CD244 on SM, and "b" signifies the effect of SM on HNC) and the product method (a × b), and (ii) the direct effect (c′ = c – a × b). The proportion mediated is calculated by dividing the indirect effect by the total effect.

Figure 2 Forest plot to visualize causal effect of each single SNP on total HNC risk.

Figure 3 Forest plot to visualize the causal effects of SM with CD244 and HNC.

Figure 4 Schematic diagram of the SM mediation effect.

[1] Nersesian S, Carter EB, Lee SN, Westhaver LP, Boudreau JE. Killer instincts: natural killer cells as multifactorial cancer immunotherapy. Front Immunol. 2023 Nov 28;14:1269614. https://doi.org/10.3389/fimmu.2023.1269614.
[2] Mathew SO, Rao KK, Kim JR, Bambard ND, Mathew PA. Functional role of human NK cell receptor 2B4 (CD244) isoforms. Eur J Immunol. 2009 Jun;39(6):1632-41. https://doi.org/10.1002/eji.200838733.
[3] Veillette A, Latour S. The SLAM family of immune-cell receptors. Curr Opin Immunol. 2003 Jun;15(3):277-85. https://doi.org/10.1016/s0952-7915(03)00041-4.
[4] Brown MH, Boles K, van der Merwe PA, Kumar V, Mathew PA, Barclay AN. 2B4, the natural killer and T cell immunoglobulin superfamily surface protein, is a ligand for CD48. J Exp Med. 1998 Dec 7;188(11):2083-90. https://doi.org/10.1084/jem.188.11.2083.
[5] Latchman Y, McKay PF, Reiser H. Identification of the 2B4 molecule as a counter-receptor for CD48. J Immunol. 1998 Dec 1;161(11):5809-12. https://doi.org/10.4049/jimmunol.161.11.5809.
[6] Chuang SS, Kim MH, Johnson LA, Albertsson P, Kitson RP, Nannmark U, et al. 2B4 stimulation of YT cells induces natural killer cell cytolytic function and invasiveness. Immunology. 2000 Jul;100(3):378-83. https://doi.org/10.1046/j.1365-2567.2000.00031.x.
[7] Mathew SO, Kumaresan PR, Lee JK, Huynh VT, Mathew PA. Mutational analysis of the human 2B4 (CD244)/CD48 interaction: Lys68 and Glu70 in the V domain of 2B4 are critical for CD48 binding and functional activation of NK cells. J Immunol. 2005 Jul 15;175(2):1005-13. https://doi.org/10.4049/jimmunol.175.2.1005.
[8] Wu Y, Kuang DM, Pan WD, Wan YL, Lao XM, Wang D, et al. Monocyte/macrophage-elicited natural killer cell dysfunction in hepatocellular carcinoma is mediated by CD48/2B4 interactions. Hepatology. 2013 Mar;57(3):1107-16. https://doi.org/10.1002/hep.26192.
[9] Pende D, Meazza R, Marcenaro S, Aricò M, Bottino C. 2B4 dysfunction in XLP1 NK cells: More than inability to control EBV infection. Clin Immunol. 2019 Jul;204:31-36. https://doi.org/10.1016/j.clim.2018.10.022.
[10] Schlaphoff V, Lunemann S, Suneetha PV, Jaroszewicz J, Grabowski J, Dietz J, et al. Dual function of the NK cell receptor 2B4 (CD244) in the regulation of HCV-specific CD8+ T cells. PLoS Pathog. 2011 May;7(5):e1002045. https://doi.org/10.1371/journal.ppat.1002045.
[11] Chow LQM. Head and Neck Cancer. N Engl J Med. 2020 Jan 2;382(1):60-72. https://doi.org/10.1056/NEJMra1715715.
[12] Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021 May;71(3):209-249. https://doi.org/10.3322/caac.21660.
[13] Rasheduzzaman M, Kulasinghe A, Dolcetti R, Kenny L, Johnson NW, Kolarich D, et al. Protein glycosylation in head and neck cancers: From diagnosis to treatment. Biochim Biophys Acta Rev Cancer. 2020 Dec;1874(2):188422. https://doi.org/10.1016/j.bbcan.2020.188422.
[14] Mehanna H, Paleri V, West CM, Nutting C. Head and neck cancer--Part 1: Epidemiology, presentation, and prevention. BMJ. 2010 Sep 20;341:c4684. https://doi.org/10.1136/bmj.c4684.
[15] Slotte JP. Biological functions of sphingomyelins. Prog Lipid Res. 2013 Oct;52(4):424-37. https://doi.org/10.1016/j.plipres.2013.05.001.
[16] Emdin CA, Khera AV, Kathiresan S. Mendelian Randomization. JAMA. 2017 Nov 21;318(19):1925-1926. https://doi.org/10.1001/jama.2017.17219.
[17] Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet. 2014 Sep 15;23(R1):R89-98. https://doi.org/10.1093/hmg/ddu328.
[18] Zhao JH, Stacey D, Eriksson N, Macdonald-Dunlop E, Hedman ÅK, Kalnapenkis A, et al. Genetics of circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat Immunol. 2023 Sep;24(9):1540-1551. https://doi.org/10.1038/s41590-023-01588-w.
[19] Kurki MI, Karjalainen J, Palta P, Sipilä TP, Kristiansson K, Donner KM, et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature. 2023 Jan;613(7944):508-518. https://doi.org/10.1038/s41586-022-05473-8.
[20] Chen Y, Lu T, Pettersson-Kymmer U, Stewart ID, Butler-Laporte G, Nakanishi T, et al. Genomic atlas of the plasma metabolome prioritizes metabolites implicated in human diseases. Nat Genet. 2023 Jan;55(1):44-53. https://doi.org/10.1038/s41588-022-01270-1.
[21] 1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, et al. A map of human genome variation from population-scale sequencing. Nature. 2010 Oct 28;467(7319):1061-73. https://doi.org/10.1038/nature09534.
[22] Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018 May 30;7:e34408. https://doi.org/10.7554/eLife.34408.
[23] Burgess S, Small DS, Thompson SG. A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res. 2017 Oct;26(5):2333-2355. https://doi.org/10.1177/0962280215597579.
[24] Bowden J, Davey Smith G, Haycock PC, Burgess S. Consistent Estimation in Mendelian Randomization with Some Invalid Instruments Using a Weighted Median Estimator. Genet Epidemiol. 2016 May;40(4):304-14. https://doi.org/10.1002/gepi.21965.
[25] Hartwig FP, Davey Smith G, Bowden J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int J Epidemiol. 2017 Dec 1;46(6):1985-1998. https://doi.org/10.1093/ije/dyx102.
[26] Yavorska OO, Burgess S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol. 2017 Dec 1;46(6):1734-1739. https://doi.org/10.1093/ije/dyx034.
[27] Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol. 2017 May;32(5):377-389. https://doi.org/10.1007/s10654-017-0255-x.
[28] Verbanck M, Chen CY, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018 May;50(5):693-698. https://doi.org/10.1038/s41588-018-0099-7.
[29] Burgess S, Butterworth A, Thompson SG. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol. 2013 Nov;37(7):658-65. https://doi.org/10.1002/gepi.21758.
[30] Zhang Y, Liu Z, Choudhury T, Cornelis MC, Liu W. Habitual coffee intake and risk for nonalcoholic fatty liver disease: a two-sample Mendelian randomization study. Eur J Nutr. 2021 Jun;60(4):1761-1767. https://doi.org/10.1007/s00394-020-02369-z.
[31] Carter AR, Sanderson E, Hammerton G, Richmond RC, Davey Smith G, Heron J, et al. Mendelian randomisation for mediation analysis: current methods and challenges for implementation. Eur J Epidemiol. 2021 May;36(5):465-478. https://doi.org/10.1007/s10654-021-00757-1.
[32] Gao P, Lu W, Hu S, Zhao K. Differentially Infiltrated Identification of Novel Diagnostic Biomarkers Associated with Immune Infiltration in Nasopharyngeal Carcinoma. Dis Markers. 2022 Nov 17;2022:3934704. https://doi.org/10.1155/2022/3934704.
[33] Tan L, Cheng D, Wen J, Huang K, Zhang Q. Identification of prognostic hypoxia-related genes signature on the tumor microenvironment in esophageal cancer. Math Biosci Eng. 2021 Sep 7;18(6):7743-7758. https://doi.org/10.3934/mbe.2021384.
[34] Chiha J, Mitchell P, Gopinath B, Burlutsky G, Plant A, Kovoor P, et al. Prediction of Coronary Artery Disease Extent and Severity Using Pulse Wave Velocity. PLoS One. 2016 Dec 22;11(12):e0168598. https://doi.org/10.1371/journal.pone.0168598.
[35] Altvater B, Landmeier S, Pscherer S, Temme J, Schweer K, Kailayangiri S, et al. 2B4 (CD244) signaling by recombinant antigen-specific chimeric receptors costimulates natural killer cell activation to leukemia and neuroblastoma cells. Clin Cancer Res. 2009 Aug 1;15(15):4857-66. https://doi.org/10.1158/1078-0432.CCR-08-2810.
[36] Agresta L, Lehn M, Lampe K, Cantrell R, Hennies C, Szabo S, et al. CD244 represents a new therapeutic target in head and neck squamous cell carcinoma. J Immunother Cancer. 2020 Mar;8(1):e000245. https://doi.org/10.1136/jitc-2019-000245.
[37] Wang H, Luo Y, Chen H, Hou H, Hu Q, Ji M. Non-Targeted Serum Lipidomics Analysis and Potential Biomarkers of Laryngeal Cancer Based on UHPLC-QTOF-MS. Metabolites. 2022 Nov 9;12(11):1087. https://doi.org/10.3390/metabo12111087.
[38] Zheng X, Hou Z, Qian Y, Zhang Y, Cui Q, Wang X, et al. Tumors evade immune cytotoxicity by altering the surface topology of NK cells. Nat Immunol. 2023 May;24(5):802-813. https://doi.org/10.1038/s41590-023-01462-9.
[39] Farhangnia P, Ghomi SM, Mollazadehghomi S, Nickho H, Akbarpour M, Delbandi AA. SLAM-family receptors come of age as a potential molecular target in cancer immunotherapy. Front Immunol. 2023 May 11;14:1174138. https://doi.org/10.3389/fimmu.2023.1174138.
[40] Jacobi J, García-Barros M, Rao S, Rotolo JA, Thompson C, Mizrachi A, et al. Targeting acid sphingomyelinase with anti-angiogenic chemotherapy. Cell Signal. 2017 Jan;29:52-61. https://doi.org/10.1016/j.cellsig.2016.09.010.
Peer-review Terminology
Identity transparency: Single anonymized
Reviewer interacts with: Editor
Details
This is an open access article under the terms of the Creative Commons Attribution License(http://creativecommons.org/licenses/by/4.0/), which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
Publication History
Received 2025-01-30
Accepted 2025-02-23
Published 2025-03-30


